Neurodata widgets
Session metadata
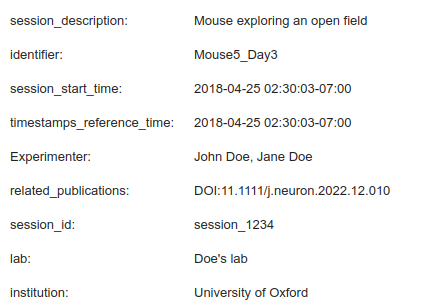
Experimental sessions metadata is stored in the pynwb.NWBFile object in NWB files and is rendered as a widget by the nwbwidgets.file.show_nwbfile function.
Subject metadata
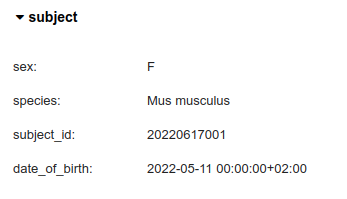
Subejct’s metadata is stored in the pynwb.file.Subject object in NWB files and is rendered as a widget by the nwbwidgets.base.show_fields function.
Extracellular electrophysiology series
Ecephys series are stored in pynwb.ecephys.ElectricalSeries objects in NWB files and are rendered as a widget by nwbwidgets.ecephys.ElectricalSeriesWidget.
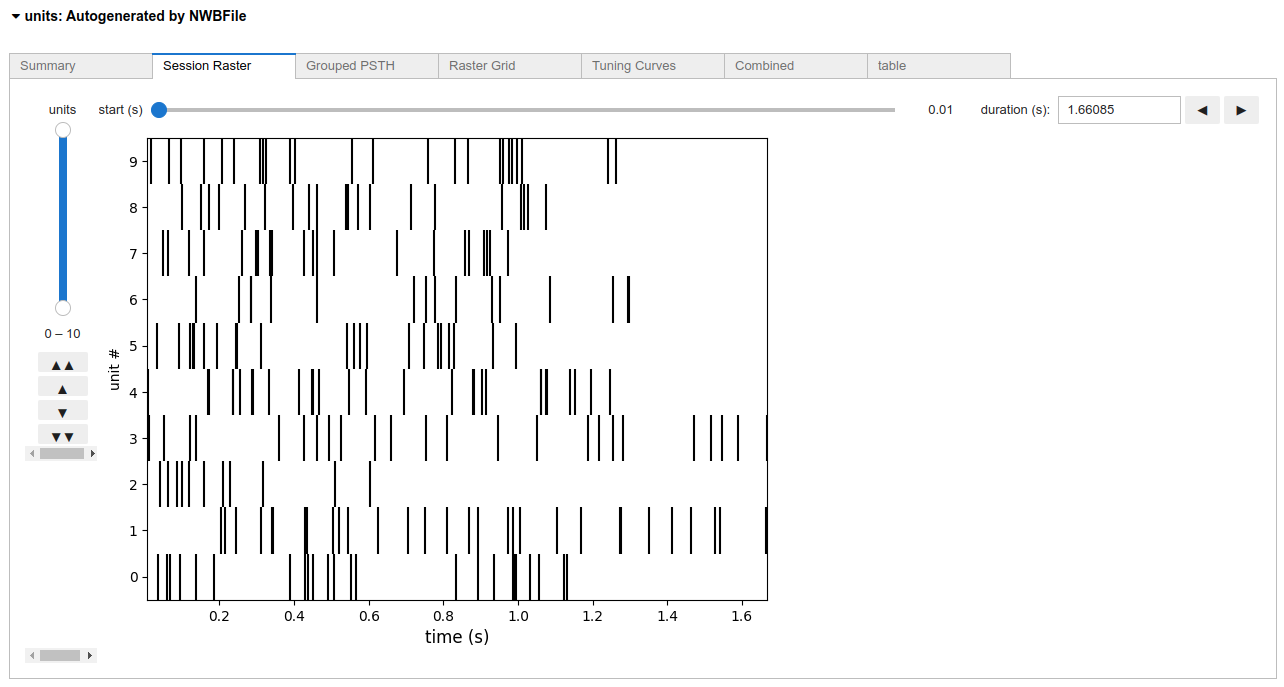
Units
Units spiking activity is stored in pynwb.misc.Units objects in NWB files and can be rendered as multiple widgets: nwbwidgets.dynamictablesummary.DynamicTableSummaryWidget, nwbwidgets.misc.RasterWidget, nwbwidgets.misc.PSTHWidget, nwbwidgets.misc.RasterGridWidget, nwbwidgets.misc.TuningCurveWidget and nwbwidgets.misc.TuningCurveExtendedWidget.
Intracellular electrophysiology series
Icephys series are stored in pynwb.icephys.PatchClampSeries and organized in pynwb.icephys.IntracellularRecordingsTable objects in NWB files. Those objects are rendered as widgets by nwbwidgets.dynamictablesummary.DynamicTableSummaryWidget, nwbwidgets.icephys.IVCurveWidget and the nwbwidgets.view.show_dynamic_table function.
Spatio-temporal behavioral series
Behavioral spatio-temporal data is stored in pynwb.behavior.SpatialSeries objects in NWB files and are rendered as a widget by the nwbwidgets.behavior.route_spatial_series function.
(EXAMPLE_GIF)
Optophysiology data
Optophysiology data can be stored as multiple objects in a NWB file: pynwb.ophys.ImageSegmentation, pynwb.ophys.TwoPhotonSeries, pynwb.ophys.PlaneSegmentation, pynwb.ophys.DfOverF and pynwb.ophys.RoiResponseSeries. Those objects are rendered as widgets by nwbwidgets.ophys.show_image_segmentation, nwbwidgets.ophys.TwoPhotonSeriesWidget, nwbwidgets.ophys.route_plane_segmentation, nwbwidgets.ophys.show_df_over_f and nwbwidgets.ophys.RoiResponseSeriesWidget.
(EXAMPLE_GIF)